

Structural genomics and modeling offer the prospect of a comprehensive knowledge of the structure of individual proteins at atomic resolution. However, extending this knowledge to (larger) protein complexes by X-ray crystallography or NMR spectroscopy has proven difficult due to technical limitations. 3D electron density maps of such complexes can be reconstructed from electron micrographs, but the resolution of these maps is typically insufficient for direct atomic modelling. A detailed understanding of the structure of macromolecular complexes therefore often requires fitting of atomic-resolution models of parts of the complex into comprehensive low-resolution density maps.
The most widely used automated fitting methods aim to maximize the cross-correlation between the EM density map and a pseudo-map computed by convolving the atomic structure with a point-spread function. The main disadvantage of this approach is its computational cost, even after Fourier acceleration of the translational search. Run times of an hour or more become prohibitive if an analysis requires multiple runs, for instance with alternative model structures. The key role of surface information in visual fitting inspired us to devise a method that maximizes surface overlap. This approach allowed the formulation of an algorithm that is typically two orders of magnitude faster than publicly available general fitting methods.
First, the template (an EM density map) and the convolved target (a structure model) are described as 3D iso-surfaces. Then, the transformations are explored that project any member of a set of test points of the target surface onto a point of the template surface and that superimposes the pair of vectors associated with these points. Finally, the 1000-2000 transformations with the highest surface overlap (fraction of superimposed voxels) are rescored using more sophisticated scores including cross-correlation.
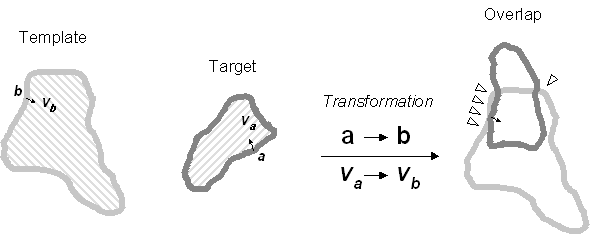
Vector-based circumference superimposition. A two-dimensional variant of the three-dimensional vector-based surface superimposition that is central to the 3SOM algorithm. For a test voxel a on the circumference of the target, the vector is calculated that approximates the normalized and inward-pointing vector that is orthogonal to the tangent line in a and with origin in a. This vector is superimposed on the vector that is associated with a voxel b on the circumference of the template. The goodness-of-fit of the transformation in question is assessed by measuring the circumference overlap (triangles). In three dimensions, a rotational degree of freedom is left around the superimposed vectors.
The (ANSI C) source code of 3SOM is freely available for all academic use, including integration in other packages, provided that adequate reference is made to the original method as described in the 3SOM paper. As 3SOM uses standard C libraries, the 3som.c file can be compiled by the command (g)cc 3som.c -o 3som -O3 -lm. Please note that we have tested 3SOM has only on a Pentium IV Linux box.
The programme consecutively asks for :
The programme produces :

 Impressum
Impressum